nnInteractive: Redefining 3D Promptable Segmentation
Highlights
- nnInteractive (2025) presents the first 3D interactive open-set segmentation model based on nnUNet, supporting a wide range of positive and negative prompts (points, scribbles, bounding boxes, and lasso) across multiple imaging modalities (CT, MR, PET, etc.).
- It is trained on an unprecedented scale with over 120 diverse 3D datasets, including a wide range of modalities, anatomical structures and label variations.
- It achieves state-of-the-art performance segmentation accuracy, interactive refinement, and clinical usability.
- Code for inference and pretrained weights are available on the official Git repo.
Introduction
3D segmentation is critical in medical imaging, biology, and industrial inspection: 3D data captures structural information that 2D images cannot. While automatic models perform well on known tasks, they fail to generalize to unseen structures or domains.
This creates a need for interactive segmentation, where users can guide and correct predictions. SAM-based models have shown great promise for 2D natural images, but extending them to 3D medical imaging remains challenging:
- 2D-only processing : most models segment slice by slice, missing volumetric context essential for anatomical consistency
- Poor generalization : models are task-specific and require retraining when data, modality, or pathology changes
- Limited interactions : only basic prompts (points, bounding boxes) are supported, no scribbles or intuitive inputs
- No accessible interface : no practical tool exists to connect users to these models efficiently
In this paper, the authors present nnInteractive, a 3D interactive segmentation framework that tackles these challenges. Rather than proposing new architectural innovations, they have 4 goals :
- Usability
- Interaction diversity
- Generalization
- Computational efficiency
nnInteractive : Network Architecture
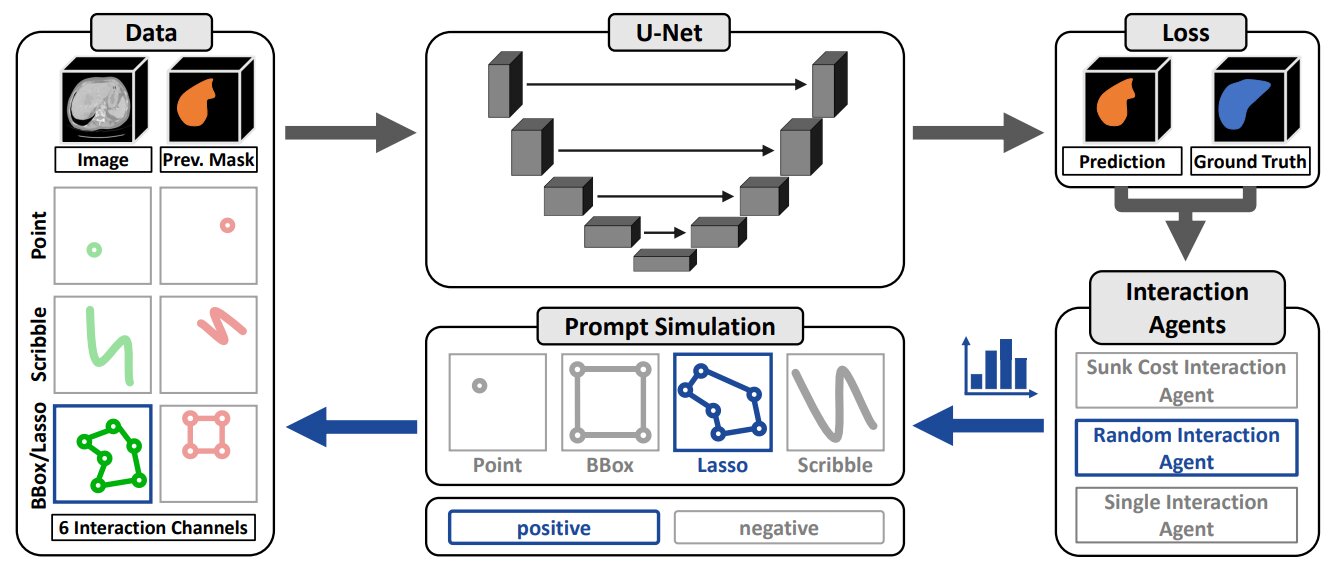
Figure 1. Overview of the nnInteractive Training Pipeline.
3D UNet remains state-of-the-art for 3D medical image segmentation despite the rise of transformer-based models in 2D. nnInteractive chooses UNet-based architecture built upon the nnU-Net framework using the Residual Encoder configuration as backbone.
Key difference from SAM: image and interaction prompts are concatenated directly as inputs, rather than fusing interactions into a latent space. This way, prompts influence the full feature extraction process from the start, enabling task-relevant learning at the highest resolution.
Interaction Input (Usability)
To minimize annotation effort, nnInteractive takes 2D prompts drawn on any plane and generates a full 3D segmentation. This is more practical than existing approaches:
- Easier than 3D annotation: clinicians only annotate on a single 2D slice, which fits naturally into standard 2D clinical workflows
- Less annotation than 2D methods : one interaction propagates to the entire volume
Each prompt type is encoded into two dedicated input channels (positive and negative).
Training
Interaction Simulation
The first prompt is sampled from the ground truth, each subsequent prompt targets the current prediction error:
- Error regions are computed as the difference between ground truth \(y\) and the prediction \(\hat{y}\)
- A random error component \(V \in \mathbb{R}^3\) is selected, with probability proportional to its size
For scribbles, lassos, and bounding boxes, a 2D slice \(S\) is sampled with probability based on the foreground volume distribution across all planes, biasing toward slices with more foreground voxels.
\(\rightarrow\) Point (Interaction diversity )
A point interaction \(p \in \mathbb{R}^3\) is sampled from the error volume \(V\) according to a probability distribution weighted by the normalized Euclidean distance from the border. Voxels near the center are favored over those near the boundary.
\(\forall x \in V : p(x) = \frac{D(x)^{\alpha}}{\sum_{z\in V}D(z)^{\alpha}}\) with \(D\) the normalized Euclidean Distance Transform (EDT) assigning higher values to voxels farther from the border (i.e., closer to the center) and \(\alpha\) controls the sampling bias (\(\alpha =8\) :centered biased approach or \(\alpha =1\): uniform sampling).
\(\rightarrow\) Bounding Box/ Lasso (Interaction diversity )
A bounding box (bbox) encloses the 2D error region in the selected slice \(S\) and is randomly augmented through three independent operations: jittering (boundary perturbed by a random offset), shifting (bbox translated by a random displacement) and scaling (random scale factor applied per dimension)
A lasso encloses the 2D error region using morphological closing and dilation, where the structuring element sizes are adapted to the local object shape via the directional Euclidean Distance Transform (EDT). The contour is further deformed by a random displacement field, sampled proportionally to the directional EDT, introducing realistic shape variability.
\(\rightarrow\) Scribble (Interaction diversity )
Inspired by ScribblePrompt 1, three types of scribbles are generated with equal probability and constant line thickness:
- Line Scribbles: connecting two random points from the 2D error mask
- Center Scribbles: extracting the skeleton of the 2D error mask, then truncating to simulate partial annotation.
- Contour Scribbles: first eroding the 2D error mask and then computing the truncated contour of the eroded object.
User behavior
To better simulate realistic user behavior across multiple interactions and avoid overfitting to unrealistic prompt strategies, three agent types are defined:
- Random agent: Selects a different prompt type at each iteration
- Sunk cost agent: Commits to one interaction type for several consecutive iterations before switching
- Single interaction agent: Uses only one interaction type throughout the entire session
Data augmentation (Generalization )
- Spacing: No resampling is performed, allowing the model to handle diverse resolutions at inference.
- Data transformation: Random scaling per dimension, transposition, and intensity inversion.
- Add pseudo labels: To improve label flexibility and reduce annotation ambiguity, pseudo-labels are incorporated during training. SAM’s automatic segment everything feature is applied to axially sampled slices to generate high-confidence supervoxels (\(\geqslant 92\%\)). These are then propagated across the remaining slices using SAM2’s video mask propagation, treating slices as sequential frames, to produce full 3D segmentations.
Auto zoom (Computational efficiency )
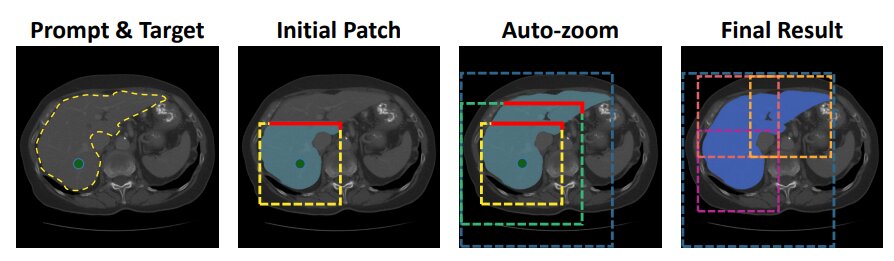
Figure 2. Auto Zoom.
Due to VRAM constraints inherent to 3D models, an adaptive autozoom strategy is introduced to reduce computational cost. The idea is to dynamically expand the region of interest (ROI) based on the predicted segmentation boundaries.
An initial crop of size \(192 \times 192 \times 192\) is centered around the first user interaction and passed through the model. The resulting prediction is checked for boundary clipping by comparison against the previous prediction (initialized as a null tensor). If the object is truncated, the ROI is expanded by a factor of 1.5, re-cropped, and downsampled to fit the model input. This process repeats until the object is fully contained within the ROI, giving a final low-resolution prediction. The prediction is then resized to the original volume size, and local patches are generated to refine the final segmentation.
This adaptive scheme minimizes computational overhead: small objects remain unaffected for faster inference, while large structures like the liver undergo progressive zoom and refinement.
Training Data (Generalization )
The model is trained on over 120 publicly available 3D segmentation datasets, comprising 64,518 volumes and 717,148 annotated objects, with \(5 \%\) held out for internal validation. The collection spans multiple imaging modalities, including Computed Tomography (CT), various Magnetic Resonance Imaging (MRI) sequences, 3D Ultrasound, Positron Emission Tomography (PET), and 3D Microscopy. At the time of publication of this paper, this data collection is unmatched in 3D medical image segmentation, exposing the model to a broad range of imaging conditions and anatomical structures.
Results
Inference Time (Computational efficiency )
The implementation is optimized for broad adoption, maintaining VRAM usage below 10 GB (under 6 GB for small objects). Small structures such as tumors and organs are segmented in 120–200 ms on an NVIDIA RTX 4090. For larger objects, inference times reaching up to 1,160 ms (\(\simeq 1.2\) s) for a liver CT and 3,700 ms (\(\simeq 3.7\) s) in rare high-resolution cases.
Comparison with SOTA methods
Evaluation is conducted on filtered datasets from the RadioActive benchmark, supplemented by four additional out-of-distribution (OOD) datasets. These datasets collectively introduce significant domain shifts in resolution, contrast, target structures, and anatomical scale, providing a rigorous test of generalization. To ensure fair comparison across state-of-the-art methods, interactions are simulated like the methods described above.
\(\rightarrow\) Qualitative comparison
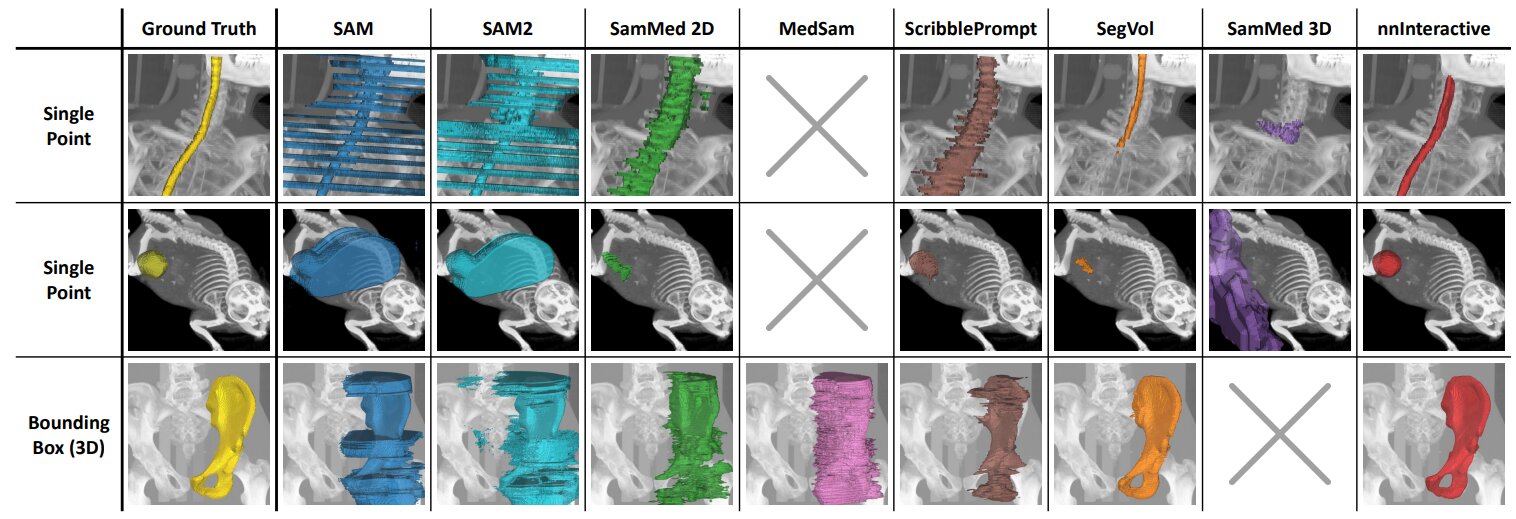
Figure 3. Qualitative comparison of interactive segmentation methods on unseen test images with different static prompting strategies: points and 3D boxes.
nnInteractive achieves the highest accuracy, closely matching the ground truth, while others struggle with precision, consistency, or volumetric adaptation.
\(\rightarrow\) Quantitative comparison
Due to the absence of competing methods capable of processing 2D bounding boxes, scribbles, and lasso in a 3D setting, all methods are compared using their only common interaction type: points.
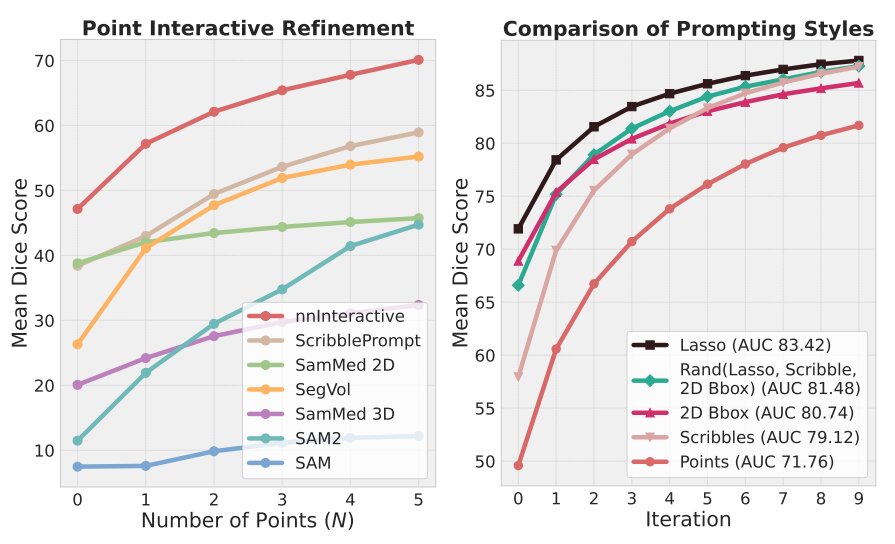
Figure 4. Interactive Performance
nnInteractive achieves the highest Dice scores in point-based refinement, with a large gap over all competitors, despite 2D models receiving N points per slice and points being nnInteractive’s weakest prompt type. Beyond points, nnInteractive’s best interaction prompts are scribbles and lasso. Interactions can be freely mixed for flexibility with better performance.
Autozoom performance
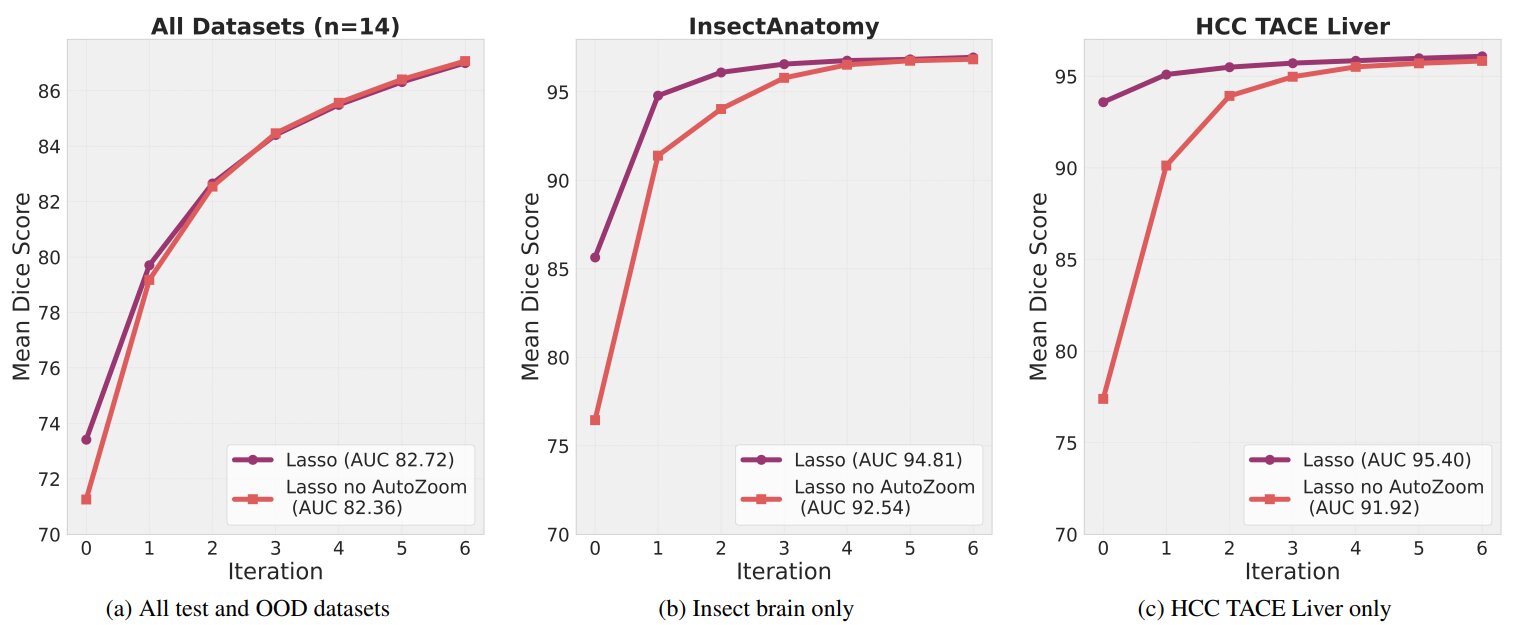
Figure 5. The effect of AutoZoom on segmentation performance
For large structures (InsectAnatomy, HCC TACE Liver datasets), AutoZoom significantly improves segmentation quality with fewer user interactions and reaching performance saturation much faster than without it (Computational efficiency ).
Ambiguity resolution
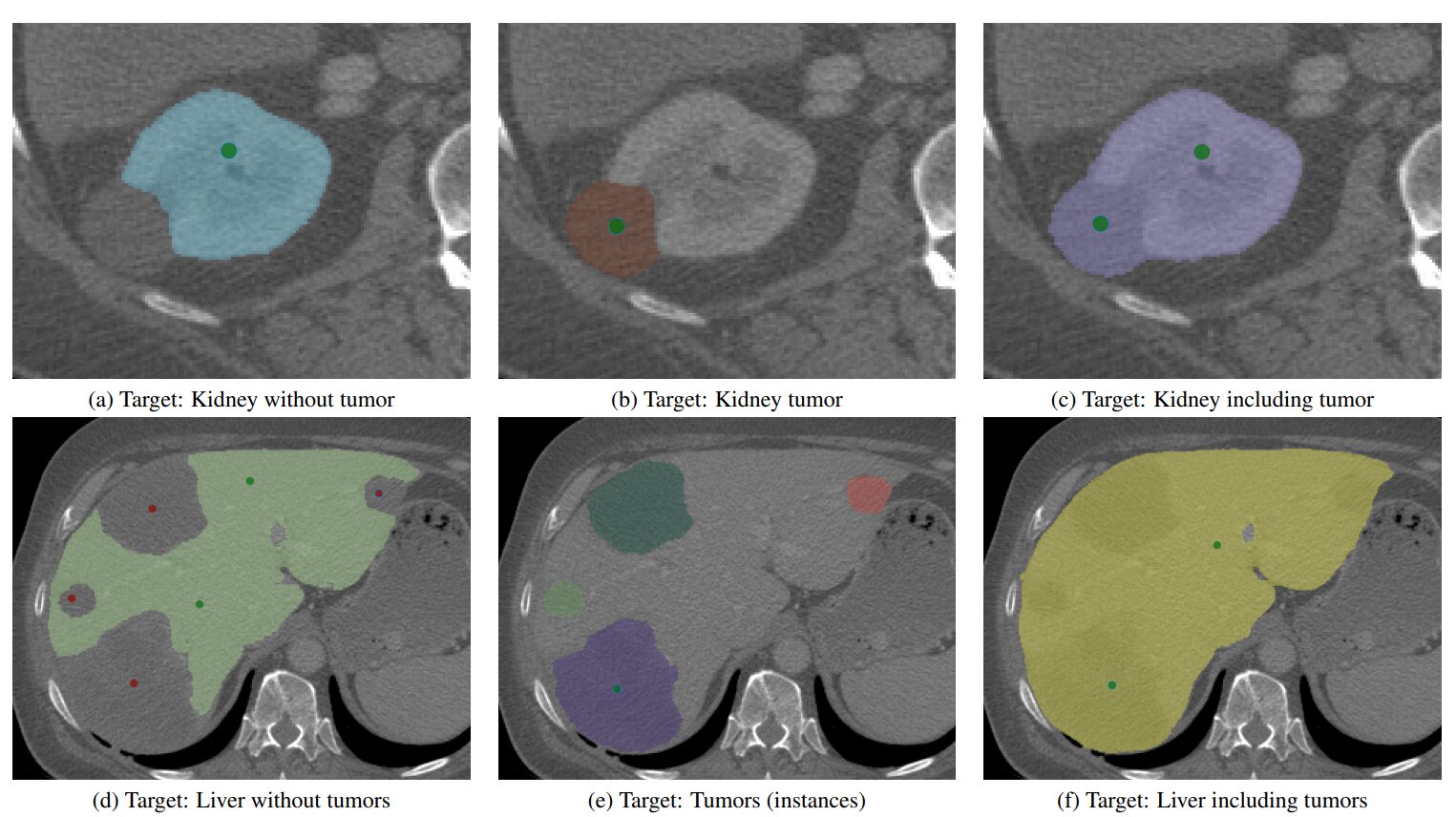
Figure 6. Image is liver 190 from the MSD Task 3 test set.
nnInteractive dynamically adapts to user input and is able to efficiently resolve ambiguities with minimal interaction (Usability).
Clinicians feedback
nnInteractive can be integrated into established imaging platforms such as Napari and MITK Workbench (Usability).
nnInteractive is benchmarked against expert manual tumor annotation in terms of accuracy and efficiency. Segmentation consistency is comparable to inter-expert agreement, with median Dice similarity scores of 0.842 (resident vs. specialist), 0.794 (specialist vs. nnInteractive), and 0.853 (resident vs. nnInteractive).
In terms of efficiency, experts completed segmentations in \(179 \pm 114\) s using nnInteractive, compared to \(635 \pm 343\) s for manual annotation (\(72\%\) reduction in time Computational efficiency ).
Conclusion
nnInteractive is a universal 3D interactive segmentation framework supporting a wide range of prompt types. It surpasses all existing methods in segmentation accuracy while significantly reducing annotation effort, and is designed for seamless adoption in both clinical and research workflows.